Evolution of mutualism
 Cooperation is common at all levels of biological organization: genes in genomes, cells in metazoan bodies, individuals in communities. Much research has been devoted to understanding the evolution of cooperation between individuals within species, where complex recognition systems, long term interactions and kin selection can all play important roles in maintaining reciprocal bonds. Less attention has been paid to mutualisms between species that lack the forms of recognition and memory characteristic of many social species.
Cooperation is common at all levels of biological organization: genes in genomes, cells in metazoan bodies, individuals in communities. Much research has been devoted to understanding the evolution of cooperation between individuals within species, where complex recognition systems, long term interactions and kin selection can all play important roles in maintaining reciprocal bonds. Less attention has been paid to mutualisms between species that lack the forms of recognition and memory characteristic of many social species.
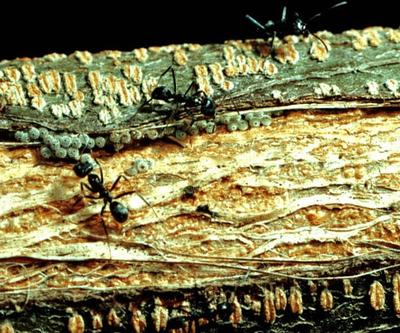 We are conducting empirical work to assess the costs and benefits of association for partners in a variety of interactions, including mutualisms between lycaenid butterflies and attendant ants, ant-plants and their inhabitants, ants and their endosymbionts and plants and their pollinators. More recently, we have also been looking at ways to apply microeconomic theory, especially the economics of information and game theory, to these well-characterized mutualisms. We are interested in why mutualism seems to persist in the face of parasites or cheaters, and whether cooperation can evolve more readily under particular environmental conditions. We are investigating factors that might promote the evolution of specialization in mutualistic interactions, and whether selection for cooperation has promoted species diversification. Approaches include analyses of the inter-specific signaling and deception, asymmetries between hosts and symbionts, adverse selection and market segmentation as applied to partner choice mechanisms, tailored models of specific mutualisms for which we have detailed behavioral data, and general models outlining the kinds of contracts that can exist between species.
We are conducting empirical work to assess the costs and benefits of association for partners in a variety of interactions, including mutualisms between lycaenid butterflies and attendant ants, ant-plants and their inhabitants, ants and their endosymbionts and plants and their pollinators. More recently, we have also been looking at ways to apply microeconomic theory, especially the economics of information and game theory, to these well-characterized mutualisms. We are interested in why mutualism seems to persist in the face of parasites or cheaters, and whether cooperation can evolve more readily under particular environmental conditions. We are investigating factors that might promote the evolution of specialization in mutualistic interactions, and whether selection for cooperation has promoted species diversification. Approaches include analyses of the inter-specific signaling and deception, asymmetries between hosts and symbionts, adverse selection and market segmentation as applied to partner choice mechanisms, tailored models of specific mutualisms for which we have detailed behavioral data, and general models outlining the kinds of contracts that can exist between species.
Phylogeny, biogeography and systematics
 We are engaged in reconstructing the phylogenies of a diversity of organisms, especially butterflies, social insects and their symbionts. These evolutionary histories are helping us to investigate how species relationships are initiated and maintained, and how they can influence rates of evolution and diversification. For example, we are interested in whether we can see the signature of mutualism on the evolutionary trajectories of interacting partners. In related museum-based work, we are using phylogenetic analysis to inform classification schemes for the systematics of different groups.
We are engaged in reconstructing the phylogenies of a diversity of organisms, especially butterflies, social insects and their symbionts. These evolutionary histories are helping us to investigate how species relationships are initiated and maintained, and how they can influence rates of evolution and diversification. For example, we are interested in whether we can see the signature of mutualism on the evolutionary trajectories of interacting partners. In related museum-based work, we are using phylogenetic analysis to inform classification schemes for the systematics of different groups.
 We established and maintain the MCZ Lepidoptera DNAs and Tissues Collection with more than 25,000 fully identified and databased specimens (mostly in the family Lycaenidae and representing all the major lineages of this group). We are currently databasing the entire pinned collection of Lepidoptera in the MCZ in order to make these specimens and their collection details readily accessible to researchers worldwide.
We established and maintain the MCZ Lepidoptera DNAs and Tissues Collection with more than 25,000 fully identified and databased specimens (mostly in the family Lycaenidae and representing all the major lineages of this group). We are currently databasing the entire pinned collection of Lepidoptera in the MCZ in order to make these specimens and their collection details readily accessible to researchers worldwide.
Functional analyses of plant–insect–pathogen interactions
 As part of our focus on species interactions, we have been exploring the functional mechanisms mediating host/ parasite interactions. In particular, we are collaborating with Fred Ausubel’s laboratory (Department of Genetics, Harvard Medical School) to develop model systems that bring together the molecular and genomic tools available for Arabidopsis with ecologically relevant pathogens and insect herbivores.
As part of our focus on species interactions, we have been exploring the functional mechanisms mediating host/ parasite interactions. In particular, we are collaborating with Fred Ausubel’s laboratory (Department of Genetics, Harvard Medical School) to develop model systems that bring together the molecular and genomic tools available for Arabidopsis with ecologically relevant pathogens and insect herbivores.

We are also working with Aravi Samuel’s laboratory (Department of Physics) to develop tracking devices for high through-put assays of behavior. The characterization of conserved signaling pathways that apply to many host-parasite interactions began with studies of plants and their natural enemies, and with Arabidopsis in particular. Although hosts in nature are typically attacked by multiple pathogens and parasites simultaneously, most studies have focused on interactions with a single natural enemy. We have developed a three-way, plant-pathogen-herbivore model system, involving Arabidopsis, Pseudomonas syringae (Ps), and various leaf-chewing insects such as moths (Trichoplusia ni), butterflies (Pieris rapae) and leaf-mining flies (species of Scaptomyza). The extensive genetic tools available for both plant and microbial pathogens in these experiments has greatly facilitated this research, and we are working now to develop similar genetic tools for a model herbivorous insect. Our long-term goal is to use field and laboratory experiments with an ecologically relevant and genetically tractable system to dissect plant–insect–pathogen interactions. We hope to elucidate both underlying molecular pathways and ecological and evolutionary consequences of these complex species interactions, which serve as both biomedical models of host/ virulence interactions, and agricultural models of host/plant defense associations.
Biodiversity and life history evolution of insects
 Discovering processes to account for patterns in the diversity of life is a major goal of organismic biology, and we have combined ecological (niche breadth) and evolutionary (phylogenetic) approaches to studying species diversity of insects. In particular, we have focused on the diversity of Lepidoptera (butterflies and moths), in the tropical forests of Southeast Asia. Many new insights have come from the efforts of the Center for Tropical Forest Science (CTFS) to inventory plant diversity in a network of tropical forest dynamics plots (FDPs) located throughout the world and to monitor this diversity over time. No comparable data are currently available for insects in these plots, and we are working with CTFS to develop an arthropod monitoring scheme.
Discovering processes to account for patterns in the diversity of life is a major goal of organismic biology, and we have combined ecological (niche breadth) and evolutionary (phylogenetic) approaches to studying species diversity of insects. In particular, we have focused on the diversity of Lepidoptera (butterflies and moths), in the tropical forests of Southeast Asia. Many new insights have come from the efforts of the Center for Tropical Forest Science (CTFS) to inventory plant diversity in a network of tropical forest dynamics plots (FDPs) located throughout the world and to monitor this diversity over time. No comparable data are currently available for insects in these plots, and we are working with CTFS to develop an arthropod monitoring scheme.
 In particular, we have been analyzing the feeding habits of lepidopteran larvae at Khao Chong, a CTFS plot in southern Thailand, and we hope to extend this analysis to other CTFS sites falling along a latitudinal gradient in Asia. This will enable us to answer questions such as: How specialized are larval Lepidoptera on plants in tropical forests, and does specialization increase towards the equator? What is the beta-diversity of Lepidoptera in SE Asian forests? Is tropical insect diversity best explained by stochastic or deterministic processes? By focusing our studies at sites established and maintained by CTFS, we are capitalizing on the wealth of plant data already available for each of the FDPs. The caterpillars of many tropical Lepidoptera are undescribed, so to identify caterpillars we have been using classical rearing techniques as well as DNA barcoding to match caterpillars and adults. Our long-term goal is to use data collected from CTFS plots to reveal patterns and processes governing the diversity of tropical insects.
In particular, we have been analyzing the feeding habits of lepidopteran larvae at Khao Chong, a CTFS plot in southern Thailand, and we hope to extend this analysis to other CTFS sites falling along a latitudinal gradient in Asia. This will enable us to answer questions such as: How specialized are larval Lepidoptera on plants in tropical forests, and does specialization increase towards the equator? What is the beta-diversity of Lepidoptera in SE Asian forests? Is tropical insect diversity best explained by stochastic or deterministic processes? By focusing our studies at sites established and maintained by CTFS, we are capitalizing on the wealth of plant data already available for each of the FDPs. The caterpillars of many tropical Lepidoptera are undescribed, so to identify caterpillars we have been using classical rearing techniques as well as DNA barcoding to match caterpillars and adults. Our long-term goal is to use data collected from CTFS plots to reveal patterns and processes governing the diversity of tropical insects.
